How random is evolution?
Often, we like to think of evolution fairly anthropomorphically; as if natural selection actively decides what is, and what isn’t, best for the evolution of a species (or population). Of course, there’s not some explicit Evolution God who decrees how a species should evolve, and in reality, evolution reflects a more probabilistic system. Traits that give a species a better chance of reproducing or surviving, and can be inherited by the offspring, will over time become more and more dominant within the species; contrastingly, traits that do the opposite will be ‘weeded out’ of the gene pool as maladaptive organisms die off or are outcompeted by more ‘fit’ individuals. The fitness value of a trait can be determined from how much the frequency of that trait varies over time.
So, if natural selection is just probabilistic, does this mean evolution is totally random? Is it just that traits are selected based on what just happens to survive and reproduce in nature, or are there more direct mechanisms involved? Well, it turns out both processes are important to some degree. But to get into it, we have to explain the difference between genetic drift and natural selection (we’re assuming here that our particular trait is genetically determined).

When we consider the genetic variation within a species to be our focal trait, we can tell that different parts of the genome might be more related with natural selection than others. This makes sense; some mutations in the genome will directly change a trait (like fur colour) which might have a selective benefit or detriment, while others might not change anything physically or change traits that are neither here-nor-there under natural selection (like nose shape in people, for example). We can distinguish between these two by talking about adaptive or neutral variation; adaptive variation has a direct link to natural selection whilst neutral variation is predominantly the product of genetic drift. Depending on our research questions, we might focus on one type of variation over the other, but both are important components of evolution as a whole.
Genetic drift
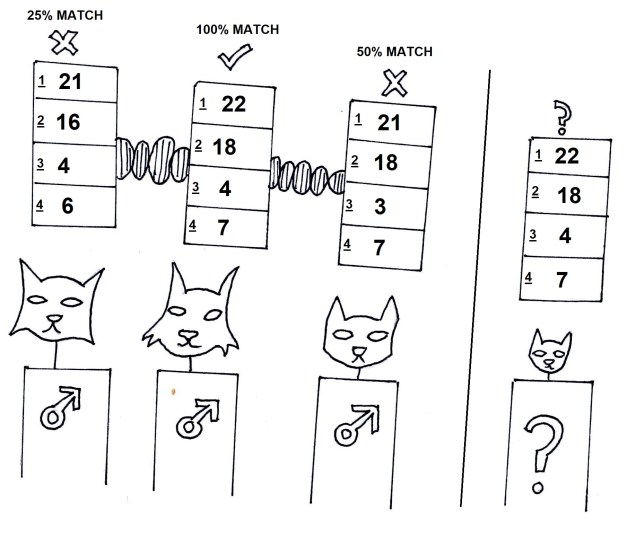
Genetic drift is considered the random, selectively ‘neutral’ changes in the frequencies of different traits (alleles) over time, due to completely random effects such as random mutations or random loss of alleles. This results in the neutral variation we can observe in the gene pool of the species. Changes in allele frequencies can happen due to entirely stochastic events. If, by chance, all of the individuals with the blue fur variant of a gene are struck by lightning and die, the blue fur allele would end up with a frequency of 0 i.e. go extinct. That’s not to say the blue fur ‘predisposed’ the individuals to be struck be lightning (we assume here, anyway), so it’s not like it was ‘targeted against’ by natural selection (see the bottom figure for this example).
Because neutral variation appears under a totally random, probabilistic model, the mathematical basis of it (such as the rate at which mutations appear) has been well documented and is the foundation of many of the statistical aspects of molecular ecology. Much of our ability to detect which genes are under selection is by seeing how much the frequencies of alleles of that gene vary from the neutral model: if one allele is way more frequent than you’d expect by random genetic drift, then you’d say that it’s likely being ‘pushed’ by something: natural selection.

Natural selection
Contrastingly to genetic drift, natural selection is when particular traits are directly favoured (or unfavoured) in the environmental context of the population; natural selection is very specific to both the actual trait and how the trait works. A trait is only selected for if it conveys some kind of fitness benefit to the individual; in evolutionary genetics terms, this means it allows the individual to have more offspring or to survive better (usually).
While this might be true for a trait in a certain environment, in another it might be irrelevant or even have the reverse effect. Let’s again consider white fur as our trait under selection. In an arctic environment, white fur might be selected for because it helps the animal to camouflage against the snow to avoid predators or catch prey (and therefore increase survivability). However, in a dense rainforest, white fur would stand out starkly against the shadowy greenery of the foliage and thus make the animal a target, making it more likely to be taken by a predator or avoided by prey (thus decreasing survivability). Thus, fitness is very context-specific.
Who wins? Drift or selection?
So, which is mightier, the pen (drift) or the sword (selection)? Well, it depends on a large number of different factors such as mutation rate, the importance of the trait under selection, and even the size of the population. This last one might seem a little different to the other two, but it’s critically important to which process governs the evolution of the species.
In very small populations, we expect genetic drift to be the stronger process. Natural selection is often comparatively weaker because small populations have less genetic variation for it to act upon; there are less choices for gene variants that might be more beneficial than others. In severe cases, many of the traits are probably very maladaptive, but there’s just no better variant to be selected for; look at the plethora of physiological problems in the cheetah for some examples.
Genetic drift, however, doesn’t really care if there’s “good” or “bad” variation, since it’s totally random. That said, it tends to be stronger in smaller populations because a small, random change in the number or frequency of alleles can have a huge effect on the overall gene pool. Let’s say you have 5 cats in your species; they’re nearly extinct, and probably have very low genetic diversity. If one cat suddenly dies, you’ve lost 20% of your species (and up to that percentage of your genetic variation). However, if you had 500 cats in your species, and one died, you’d lose only <0.2% of your genetic variation and the gene pool would barely even notice. The same applies to random mutations, or if one unlucky cat doesn’t get to breed because it can’t find a mate, or any other random, non-selective reason. One way we can think of this is as ‘random error’ with evolution; even a perfectly adapted organism might not pass on its genes if it is really unlucky. A bigger sample size (i.e. more individuals) means this will have less impact on the total dataset (i.e. the species), though.

Both genetic drift and natural selection are important components of evolution, and together shape the overall patterns of evolution for any given species on the planet. The two processes can even feed into one another; random mutations (drift) might become the genetic basis of new selective traits (natural selection) if the environment changes to suit the new variation. Therefore, to ignore one in favour of the other would fail to capture the full breadth of the processes which ultimately shape and determine the evolution of all species on Earth, and thus the formation of the diversity of life.