Gene flow and introgression
Genetic variation remains a key component of not only understanding the process and history of evolution, but also for allowing evolution to continue into the future. This is the basis of the concept of ‘evolutionary potential’ – the available variation within a population or species which may enable them to adapt to new environmental stressors as they occur. With the looming threat of contemporary climate change and environmental transformations by humanity, predicting and supporting evolutionary potential across the diversity of life is critical for conserving the stability of our biosphere.
One aspect of conservation management that we’ve discussed on several occasions is the idea of genetic rescue: supplementing the genetic diversity of particularly depauperate or threatened populations with alleles from another (more robust) population through captive breeding or reintroduction. The basis of this process is that increasing the genetic diversity of a population contributes to their adaptive potential by allowing natural selection to act on a greater diversity of traits. Without the added diversity, no traits present in the population may be adaptive enough to respond to a particular threat and the population is at risk of extinction.

Pros and cons: adaptive or maladaptive?
However, adding alleles via gene flow in genetic rescue experiments could lead to two different types of outcomes, based on the success and function of the genetic diversity being added. If the alleles added into the population are positive and enable the population to respond to a new stressor (i.e. are under positive selection), then we would consider this successful genetic rescue as it has increased the adaptive potential of the population. Contrastingly, if the allele confers a negative (maladaptive) trait, natural selection will act against it and these detrimental alleles will cause a reduction of fitness in the population. If this allele is widespread across the population enough, this could severely damage the overall survivability of the population and result in outbreeding depression. We’ve discussed the balance of these two factors in more detail here.
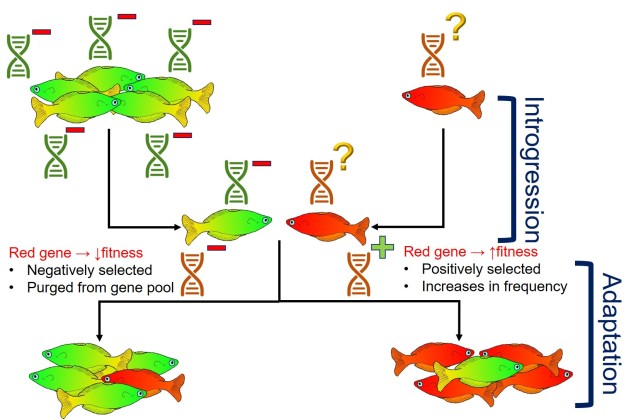
Generally, genetic rescue seems to be broadly effective. Natural selection tends to purge maladaptive alleles from the population fairly rapidly after introduction (especially if they are extremely poorly adapted) and in some cases the circumstances are so dire that it would be difficult to decrease the fitness of the population. That said, there have been examples of ‘genetic rescue gone wrong’, where introduction of new genetic variation has had a notable negative impact on the stability of the population. This cases are extremely rare, but it’s important to determine the most effective way to do translocations and genetic rescue before introductions occur.
In nature
For the most part, when we’ve discussed the potential for gene flow to allow for adaptation to occur we’ve been considering experimental or man-made scenarios. One might argue that, to some degree, this is defying nature by bringing together populations and combinations of alleles that never existed in the past (indeed, this is kind of the point). But is it wholly an anthropogenic phenomenon? Gene flow and introgression undoubtedly occurs in the wild, so does it lead to an increase in adaptive potential often? How can we tell? Let’s take a look at a few cool case studies.
Study 1: the ‘gray’ of the wolf
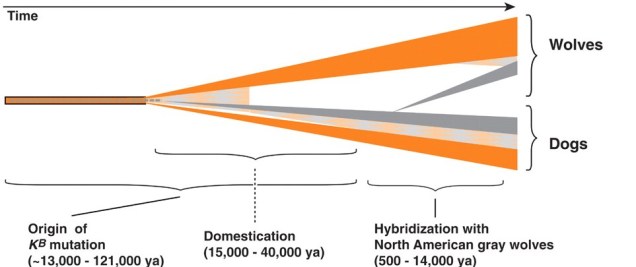
The North American gray wolf is renowned for its impressive stature and iconic coat colour. But did you know that this colour isn’t originally a wolf trait, and actually the result of adaptive introgression? In a thorough genomic study, Anderson et al. (2009) demonstrated that the genetic variant responsible for melanism in young wolves – the K allele of the gene Melanocortin 1 receptor (Mc1r) – did not originally evolve within the wolf species. Instead, the K allele originated within breeds of domestic dogs as selective breeding drove the ‘evolution’ of new traits. When the domestic dog was introduced into the Americas, likely following recolonization across the Bering Strait with the migration of humans 12 – 14 thousand years ago, hybridisation of gray wolves and these domestic dogs lead to gene flow. The K allele was introgressed across from dogs into the wolves: since the allele codes for a melanistic coat colour, the resultant hybrids were darker in fur.
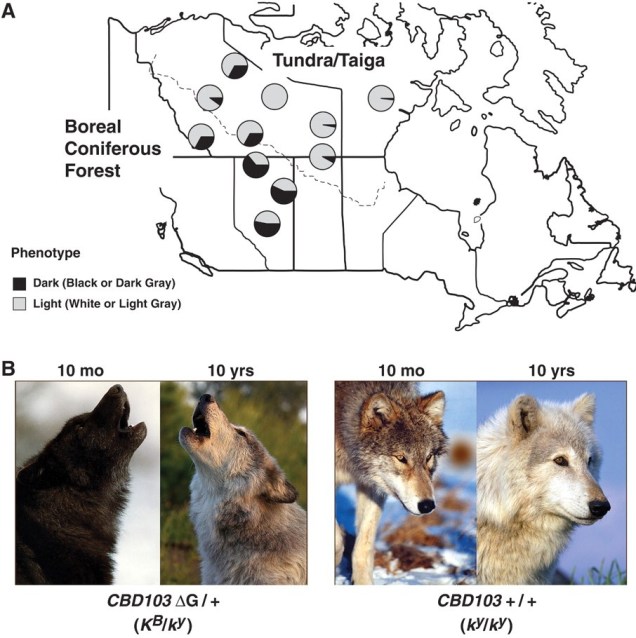
Darker fur can be an adaptive trait under dimly-lit forested habitats, and thus was positively selected under natural selection. Years later, and the darker fur trait has spread widely across populations of grey wolves inhabiting darker environments. Anderson et al. suggested that this same pattern and pathway was similarly responsible for the adaptation of melanism in coyotes and Italian gray wolves, highlighting the surprising adaptive potential added through human introduction of a domestic species.
Study 2: On coloured wings
A similar study by Pardo-Diaz et al. (2012) showed a similar history of adaptive introgression of a colour-associated gene, albeit this time in Heliconius butterflies. These butterfly species are often considered a key model in adaptive radiation given their variety of colours and wing shapes, in combination with systems of mimicry in which very different species appear to converge onto similar phenotypes. This multi-faceted complex of factors provides an interesting scenario for testing different processes of evolution. In their study, Pardo-Diaz et al. focussed on the formation of red-coloured bands on the wings of many different species of Heliconius butterflies to try and understand under which processes these traits evolved.

Remarkably, they found that much of the mimicked red-colour banding across the different species originated as a case of adaptive introgression. Transferral of the HmB453k marker (associated with the optix gene) from H. melpomene into H. timareta drove the formation of the same red-banding in the latter. The result of this is implicit mimicry – with the two species looking similar to one another – and under the process of Müllerian mimicry the trait evolved widely in H. timareta. Thus, evolution of a mimicry trait was facilitated by adaptive introgression from its target. Neat!
Study 3: (Not so) close to home
But maybe these examples are a little too detached for you. Lucky for you, even within humans there are clear examples of adaptive introgression. You may be aware that, prior to their extinction, Neandarthals and modern humans regularly interbred and hybridised. In fact, because of this, some people may owe as much as 4% of their genome to Neandarthal ancestors! This is a non-trivial amount of genetic variation (I’ve heard the comparison that, by volume, more ‘Neandarthal’ DNA exists now than at any point in the past…): could this be used for adaptation?
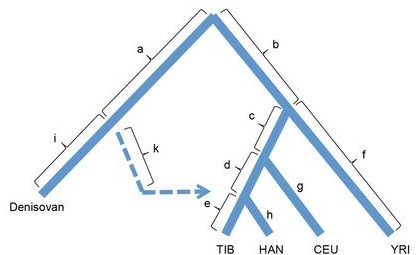
The short answer is: yes! Past hybridisation of extinct hominids and modern humans resulted in the introgression of a number of different alleles and loci, and among them was an allele for the gene EPAS1. This gene plays an important role in hypoxia pathways and has been previously shown to be under extremely strong selection in modern humans occupying high elevations – most notably, those in the Himalayas and the Tibetan Plateau. Thorough genomic interrogation by Huerta-Sanchéz et al. (2014) demonstrated that the most plausible origination of this gene was from adaptive introgression with Denisovans (another extinct species of humans). Without this hybridisation, modern humans may never have colonised some of the heights we see today.
The role of adaptive introgression
Together, these studies demonstrate some of the ways in which introgression – either naturally occurring or through human intervention – may actually be critically important in driving evolution. As environments across the globe become increasingly stressful and unstable, adaptive introgression and genetic rescue presents an avenue for supplementing the adaptive potential of many threatened populations and species.

4 thoughts on “What’s yours is mine: evolution by adaptive introgression”