Overview of 2020
As you may have gathered, The G-CAT has been significantly less active in this our most Cursed year. There are a number of reasons for that – not just the overall disaster that has been world events – including the fact that this was the last year of my PhD. I’m delighted to announce that now, after ~3.5 years of hard work, I am officially Dr. Buckley (not Dr. G-CAT, as I may have led you to believe)!

I’ve talked about my thesis in some aspect over the years, and certainly my research focus has filtered through the various topics discussed in this blog. But I’ve never really provided a (succinct) overview of the thesis itself and some of the key findings. So without further ado, let’s take a (relatively) quick look at what I studied. If you would like to know more about my work, you can find my full and publicly available thesis here.
A summary of my thesis
Through my thesis, I aimed to understand the complex interaction of environmental and evolutionary history in a group of endangered freshwater fish – the pygmy perches. Although this is important work for their conservation, the broader aim was to use these cute little fish as a ‘model’ for understanding more broadly how environmental changes over the last few millions of years have impacted the evolution of species in temperate Australia. I used genomic data from species spread across southeast and southwest Australia, combined with some complex environmental and statistical modelling approaches. The findings from this thesis were expected to inform the conservation management of not only pygmy perches but general conservation practices and evolutionary understanding of these regions.

Chapter 1: Phylogeography of southern pygmy perch
The first (data) chapter of my thesis focused on the evolutionary history of one of our friends here on The G-CAT: the southern pygmy perch. An endangered freshwater fish endemic to southeast Australia, the southern pygmy perch is threatened by a combination of human modification of aquatic habitats as well as the looming impact of contemporary climate change. However, we don’t know a lot about how they likely responded to some significant environmental changes in the past, which may provide some insight into how they could be affected in the near future.
For this chapter, I used a combination of genomic data collected from over 100 southern pygmy perch spread across 24 populations, contemporary climate data, historical climate data (ranging back to the Pliocene, over 3 million years ago!) and some rather complex coalescent modelling. There was a lot of analyses worked into this chapter, and rather than go through it all I will just link to the pre-print on bioRxiv (the paper has changed somewhat since it was uploaded). Currently, this chapter is undergoing revisions for submission to Molecular Ecology and will hopefully be published in the coming year.

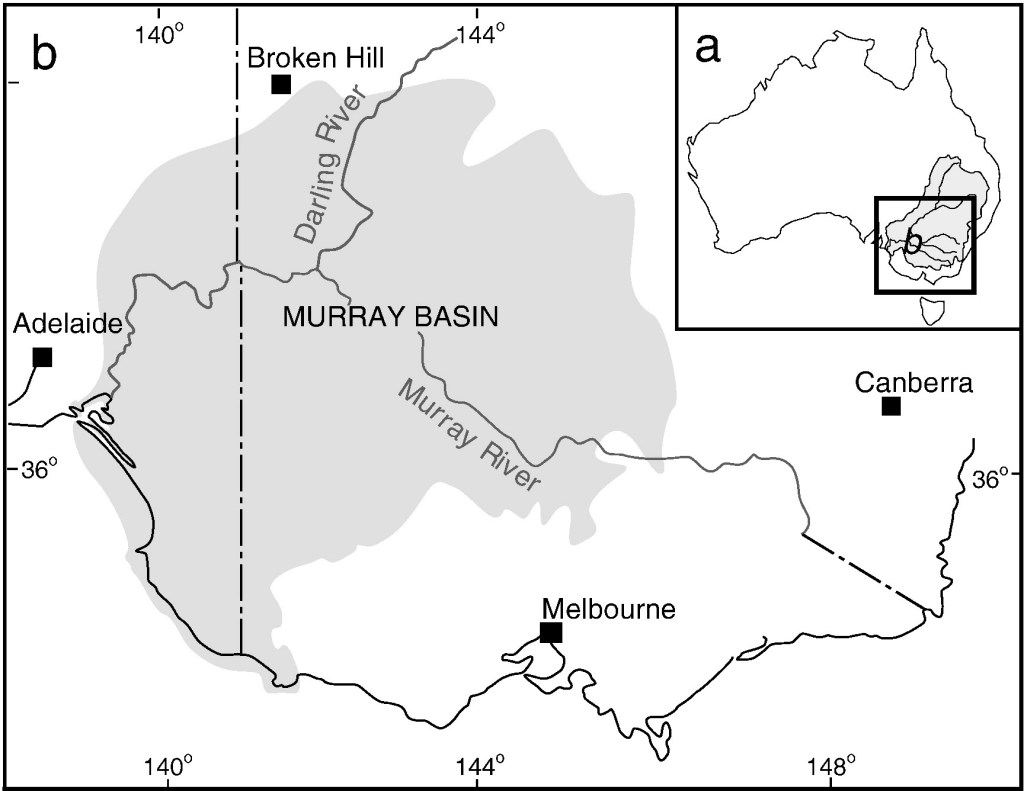
Overall, I found that major environmental changes associated with sea levels and aridification had significant impacts on the overall evolution and divergence of populations, including the formation of a cryptic species dubbed Nannoperca ‘flindersi’. One such change was the formation of a historical and massive lake within the Murray-Darling Basin named Lake Bungunnia. During its existence (from ~3 million years ago), Lake Bungunnia acted as a conduit for southern pygmy perch to colonise the inland from a coastal origin, which now makes up a major portion of their distribution. However, the demise of the lake ~700 thousand years ago, this connection was lost and now there is strong genetic differentiation between inland and coastal lineages of southern pygmy perch. There are a number of other localised environmental changes which also appear to have impacted divergence across the species, but the key finding here is the interactive effect of both sea level change and aridification on isolating populations – both of which are likely to be exacerbated under future climate change.
Chapter 2: Comparative phylogeography of southern and Yarra pygmy perches
I then expanded on these results in my next chapter by comparing evolutionary (especially demographic) histories of southern and Yarra pygmy perch. Although similar in many ways (including body shape, diet, habitat requirements), Yarra pygmy perch are even more threatened than southern pygmy perch because of extremely low genetic diversity, narrower distribution and lack of recovery from drought conditions during the turn of the century. One of the main aim of this chapter was to evaluate what role evolutionary history played in these differences – were Yarra pygmy perch simply (evolutionarily) pre-disposed to handle environmental change worse?

I focused on co-occurring populations of the species across coastal southeast Australia (to try to broadly control for environment). I compared phylogenetic, distribution and demographic histories of the two species, with a particular focus on coalescent demographic modelling both independently for each species and within a comparative approach. Interestingly, there were some key similarities and differences between the two. The most dramatic difference was in their distributions over time – during the Pleistocene, southern pygmy perch showed a dramatic expansion as more suitable habitat became available across the region. Contrastingly, Yarra pygmy perch appear to never have expanded and maintained a relatively stable distribution over millions of years. The next question was “why”?
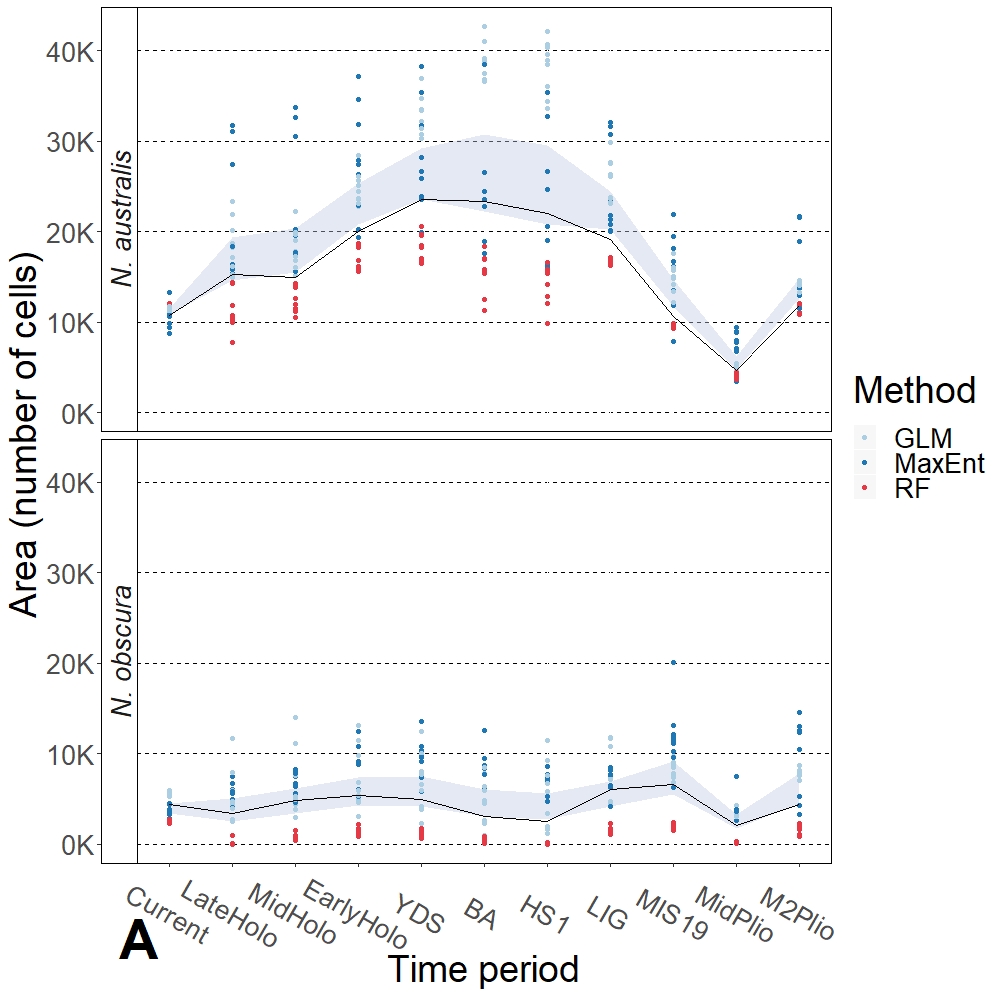
One relevant factor might be that the two species show very different levels of genetic diversity, and I proposed that the interaction of genetic diversity and the selective pressure at the margins of the distribution drives this discordance. Even historically, southern pygmy perch likely had higher genetic diversity which may have allowed them to adapt and expand into new habitat as it became available in the Pleistocene. In contrast, Yarra pygmy perch appear to have always had low genetic diversity (this is not a recent phenomenon) which might have meant that they lacked the adaptive potential to similarly expand despite their ecological similarities. This paints a bleak picture for our understanding of their ability to respond to future climate change and highlights the importance of genetic-based conservation work for the species.
Chapter 3: Phylogeography of western pygmy perches
The last data chapter of my thesis focused on pygmy perch species on the opposite side of the country – the western and little pygmy perches in southwest western Australia. Previous work from others and my Honours have suggested that western pygmy perch is actually comprised of multiple cryptic species (more on that here), which might be connected with the region’s identity as one of the only globally recognised biodiversity hotspots in Australia.
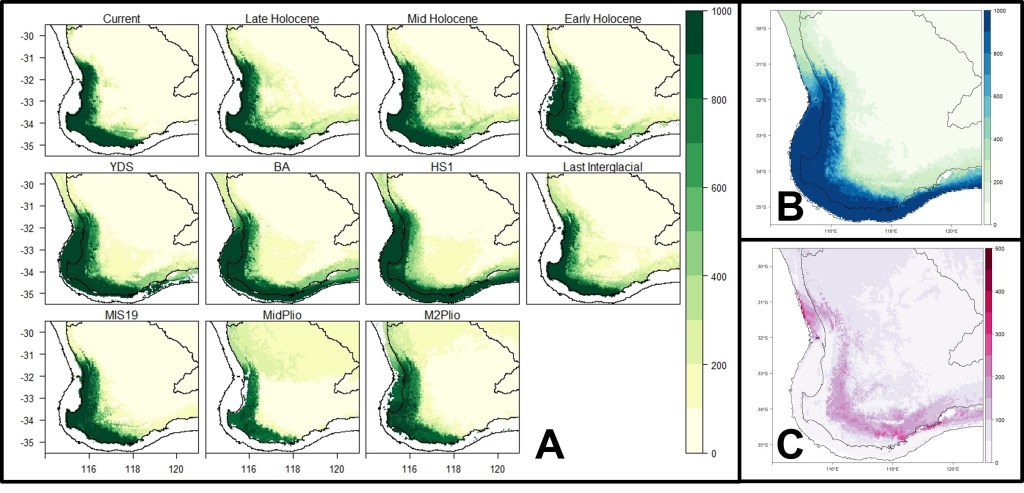
Similar to the other chapters, I used genomic data from 9 different populations of the two species across the region in several different approaches, including in conjunction with contemporary and historical climatic data. The results revealed very strong differentiation between populations of the two species (even greater than in the other species), which conservatively included at least two genetically differentiated species of western pygmy perch. These species diverged a very long time ago (~9 million years ago) and showed very little genetic exchange since. This was supported by a relatively widespread and generally stable distribution since the Pliocene, suggesting that these lineages were maintained in isolation by the stability of the environment (instead of rapid changes causing localised extinctions or reconnecting populations as in the southeast).
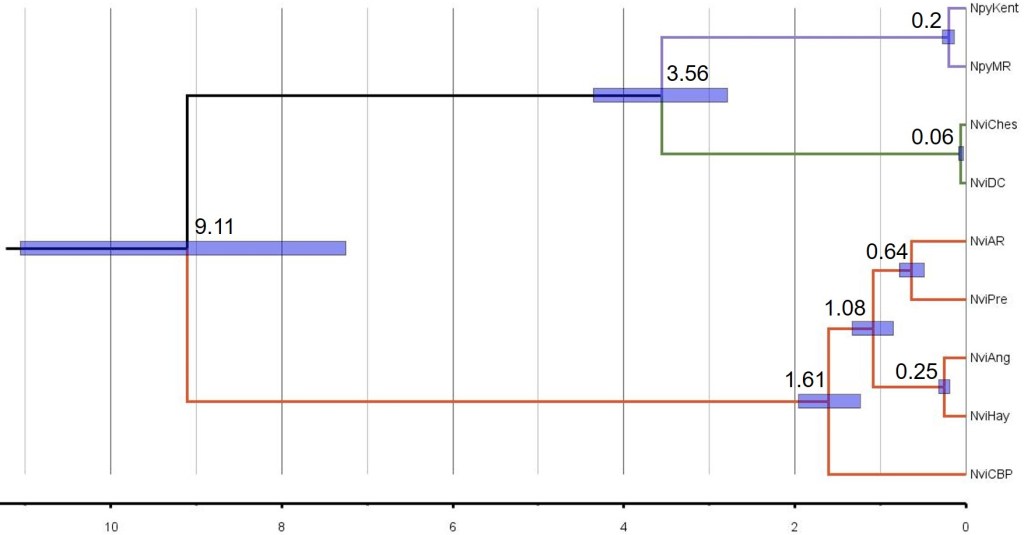
I expanded on some of this work by looking at how the most differentiation sections of the genome (‘fixed differences’) related to particular molecular functions by comparing them to the annotated genome of the southern pygmy perch (a very recent development and as yet unpublished). This aspect is a little complicated, and I will likely expand on the process more in the future. But interestingly, a number of these highly differentiated regions related to the way that chromosomes are arranged during cell duplication – a factor which has been shown to cause reproductive isolation in crosses of different species of fruit flies. Although not unequivocal evidence for their identity as true species, this finding suggests that internal molecular mechanisms might play a role in the isolation of (and lack of gene flow between) these cryptic species. Hopefully in the future I will be able to expand on this approach to provide more robust evidence for reproductive isolation, and test it with some other species.
What’s next?
All in all, I’m proud of my thesis. It’s been a bit of an emotional rollercoaster and certainly wasn’t without it’s (many) challenges, but it’s my hope that these findings will have provide some interesting insights into the evolutionary history of Australian freshwater species and have some application for the ongoing conservation management of pygmy perches. Whether I will be able to continue this work in the future is still a standing question, but you rest assured that I will be publishing this work in journals in the near future.
Thankyou for standing by while I close this important chapter of my life, and here’s to a brighter future!

Congratulations on your Doctorate ! I do not presume to understand your work but the suggested outcomes have a familiar ring.
Granpa Pope
LikeLiked by 1 person
Thankyou Grandpa! Hope the post helped you understand some of it a little better!
LikeLiked by 1 person