Beyond mutations in the genome
Although genetic variation is, in itself, often considered to be one of the fundamental underpinnings of adaptation by natural selection, it can appear through a number of different forms. Typically, we think of genetic variation in terms of individual mutations at a single site (referred to as ‘single nucleotide polymorphisms’, or SNPs), which may vary in frequency across a population or species in response to selective pressures. However, we’ve also discussed some other types of genetic-related variation within The G-CAT before, such as differential gene expression or epigenetic markers.
Structural variation
In this post, we’re going to take a look at another type of genetic marker that is rife throughout the genome: structural variations. In short, structural variation refers to parts of the physical genome which have been structurally modified through one means or another. For example, this might include the physical relocation of particularly genes within the genome (say, movement across different chromosomes) or rearrangements of individual genes. As might be expected, this can have profound outcomes on the underpinning genetic structure that drives many phenotypes and can capture large swathes of variation within the genome. Naturally, this is particularly relevant for a number of genetic disorders, but may also relate to a swathe of non-disease-associated traits.
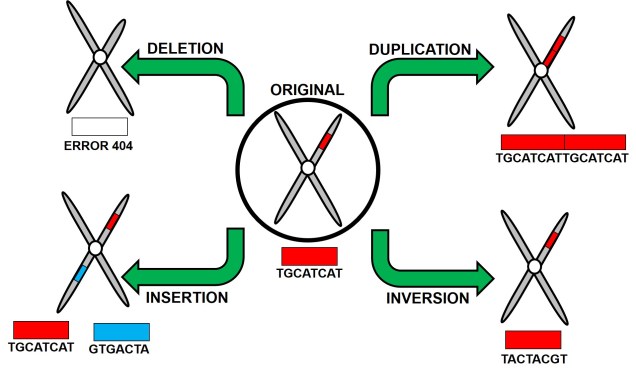
Structural variation is a somewhat umbrella term which encompasses a number of different potential components. Most commonly, structural variations refer to major rearrangements of genomic sequence through various alterations to the normal biochemical process of DNA replication (e.g. during meiosis). At their largest scale, structural variations can involve duplication of entire genes or chromosomes, or even the entire genome itself (in the case of polyploidy). However, structural variations are normally much smaller and can span 100s to 1000s of nucleotides in length. In fact, because of the sheer size of the segments that can be impacted by variations in genome structure, structural variants may actually encompass three times as much sequences as SNPs, suggesting their importance in underpinning adaptation. So, without further ado, let’s take a (brief) look at some of the types of variations and how they may impact on evolution.
Duplication
One type of structural variation that has, in some form, been touched on here before is the gene duplications. In previous posts, we’ve discussed this in the context of paralogous genes, where duplication events of a single gene lead to two separate genes with (moreorless) identical underlying sequence. Each of these genes may be related to different functions and evolve under different conditions, driving new adaptations. For many phylogenetic studies, paralogous genes are somewhat of a nuisance aspect as they can confound relative analysis if genes that appear to be the same (by sequence) but are not (paralogous) by evolutionary process.

However, this aspect belies something else rather important for evolutionary theory: that duplicated genes may take on new roles and functions as a basis of adaptation. Understandably, when duplications occur at the whole-genome scale, this can provide an extreme level of new genetic variability within the genome of a single individual. Mixing of the different (polyploid) chromosomes during meiosis can allow recombination to form new combinations of different genes that did not previously occur on the same chromosome, potentially leading to new adaptive haplotypes. This process can even drive incredibly rapid speciation if new polyploid individuals are reproductively incompatible at the genetic level with non-polyploid conspecifics.
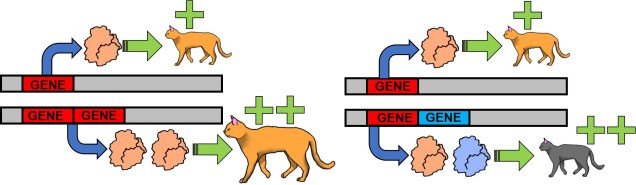
Even at smaller scales, however, duplications of genes can lead to highly adaptive phenotypes. For example, duplicated genes may cause the product of the duplicated gene – the protein itself – to also be produced in double the quantity, rapidly boosting gene expression. This increased level of available protein may be adaptive with the right gene and environment, such as HXT genes improving glucose transports into cells of yeast when duplicated.
Inversion
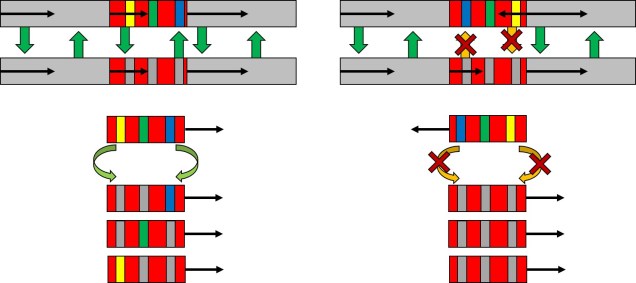
Another type of structural variation is inversions, whereby genes (or sections within genes) are rearranged in reverse order. This can occur when a chromosome breaks, separating a section of the genome from the rest of the chromosome. When the enzymatic repair system acts to fix the break, it can sometimes insert this excised section in back-to-front, causing the sequence to become inverted (relative to the rest of the chromosome). Where this occurs within the chromosome (i.e. whether it occurs across the centromere or not) dictates what type of inversion this is called: paracentric if it does not cross the centromere, or pericentric if it does.

Similar to duplication events, these altered chromosomal sequences may confer new phenotypes for natural selection to act upon. Similarly, the extent of inversions might also drive speciation if the inversion prevents recombination of chromosomes between individuals (i.e. successful reproduction, genetically). Particularly, however, inversions are great are preventing recombination even in reproducing organisms to the extent that they ‘protect’ co-adapted gene complexes from becoming lost due to mixing.
Inversions have been implicated in a number of cases of adaptive changes and speciation, ranging from plants to insects, birds and fish.
The many others
Naturally, there are a number of other structural variations which may significantly impact on the trajectory and outcomes of evolution in nature. These can include translocated genes, whereby individual genes or sections of the genome are moved from one chromosomal position to another (or even from across species in the case of horizontal transfer of transposable elements!). Alternatively, sections of the genome may be excised out through large-scale deletions. Disentangling each of these, and what role they may potentially play in adaptation, might provide interesting insights into the mechanisms and outcomes of evolution.
Like some other aspects of evolutionary genetics, such as epigenetics, we are only beginning to scratch the surface of understanding how structural variation may drive evolution. Further research into how these different chromosomal rearrangements might impact the adaptability – or not – of species will inevitably become a key feature of the future of evolutionary genetics.
I’d like to particularly thank Associate Professor Maren Wellenreuther for providing the inspiration for this week’s post, after her visit to Flinders University last week. Her research into the role of structural variation in evolution is incredibly fascinating, and I highly recommend digging into some of her work to find out more about it!
