The fundamentals of population genetics
Many times in the past, we’ve discussed the importance of genetic diversity within populations as a foundation for adaptation and evolution. It includes both adaptive variation (which encompasses genetic variation directly under natural selection), as well as neutral variation (which is predominantly generated and maintained by non-selective forces such as demographic history and genetic drift). This pool of genetic variation acts as the underlying architecture for evolution by natural selection, and is a critically important component for future and ongoing evolution.
This all sounds important from an academic perspective: that population genetics can reveal a significant amount of information about the processes and outcomes of evolution and provide novel insights into concepts that have been around for ages. But how can this information be applied to real scenarios? With the ever-growing availability of massive genetic datasets for an increasing number of species, the sheer volume of information in existence that can be used is monumental.
Of course, this is not a new topic (at all!) within this blog. Indeed, every second post on here appears to be discussing the application of genetic data in some facet or another. This time, however, I’d like to delve a little more into some specific examples of real-world application of population genetics, particularly for the direct management of natural resources. While this might sound like just more conservation management – and believe me, it definitely does include that – we can also apply our knowledge of population genetics to other more industrial faculties. For example, we might look at how genetic information is being applied in the development of more sustainable food management to deal with the imminent food crisis in the wake of overpopulation.
Although these two delineations might some very different – or even entirely contradictory to one another – they share much of the same approaches and goals (long-term sustainability of natural resources). Let’s compare some of the most common applications of genetics in conservation management and natural resource-based industry and see how they relate.
In conservation management
To start, let’s recap the application of genetics within conservation management. While this is a topic we’ve touched on a number of times within The G-CAT, there are a few distinct applications that are widely applied to conservation efforts across a swathe of species. Particularly, genetic techniques aim to provide important information about the state of population and evolutionary dynamics, which can be used to inform best management practices for the future. These fit within a wider scheme of maximising the success and output of conservation efforts, particularly when resources (namely money and staff) are often incredibly limited for many species. In this regard, conservation genetics helps to provide longer term solutions to current and imminent issues in the conservation and sustainability of species.
So without further ado, let’s take a look at some examples of population genetic applications to conservation management.
Identifying population units
One central theme of population genetics is, understandably, the definition of a population. Given the general hierarchy and transient nature of evolution (e.g. individuals within populations within species, and so on), defining what a population is as a unit can be a little tricky. From a genetic point of view, a population is a single intermixing unit of individuals which share genetic variants readily amongst one another. In this sense, populations are often considered one of the most fundamental units of evolutionary processes, as population-level shifts in allele frequencies often drive microevolution.
Within direct management, a hierarchy of populations is often important to define in terms of what spatial (and potentially temporal scale) to enact conservation strategies. To this end, we often use terms such as ‘evolutionarily significant units’ (ESUs) and ‘management units’ (MUs). Specifically, ESUs refer to population units which likely have divergent evolutionary histories (i.e. have been isolated for a significant period of time and have very different allele frequencies), whereas MUs are often not as divergent (and may result from more recent isolation of populations). Sometimes ESUs may also be colloquially referred to as ‘subspecies’ or ‘races’ to highlight their uniqueness.
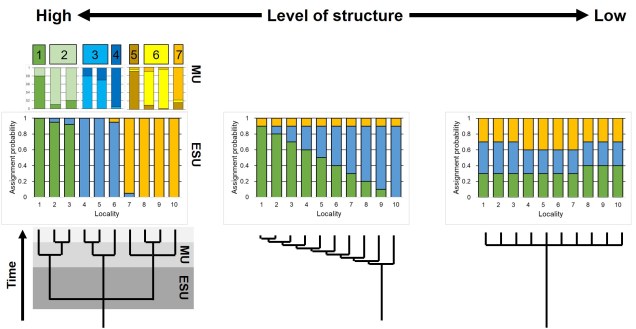
Population definitions can be applied to conservation efforts in a few different ways. This might mean limiting captive breeding and reintroductions of individuals to within ESUs to avoid the potential of outbreeding depression by ‘diluting out’ locally adaptive alleles. Contrastingly, admixture across traditionally separated populations may help to provide new (adaptive) alleles to populations at risk through genetic rescue. In any case, the concept of a population based on genetic information is critically important to effective conservation management.
Species identification and taxonomy
At a higher taxonomic level, genetic analyses may also be important for defining new species. This can be particularly important for species which are not morphologically recognisable (dubbed ‘cryptic species’) but demonstrate unique evolutionary histories and ecological functions. New species can be robustly identified from genomic sequences using methods such as species delimitation, which can statistically evaluate the likelihood of individuals belonging to a new, reproductively isolated species.

For conservation management, legislative policies are often focused at the species (or sometimes the ESU) level, with management strategies within the species considering population structure. Thus, accurately describing and denoting species is important for the development of legal processes in their protection, and may also contribute to other broad-scale conservation protocols (such as in the identification of biodiversity hotspots, which hold a disproportionate amount of biodiversity are incredibly important to conserve).
Identifying at-risk populations
With the looming (current) threat of anthropogenic climate change, it is also paramount to predict susceptibility of certain species (or populations) to changing climatic conditions. Similarly, this can best be identified with adaptive genomics methods, which can identify regions of the genome under strong ‘natural’ selection. The presence – or absence – of particular adaptive alleles may confer some resistance to the impacts of climate change, and predicting extinction risk of species is important for proactive management techniques.
In industry management
These practices might seem very specific for conservation managers – and indeed, many of them are – but they also have broader applications to natural resource industries as well. Let’s take a look at each of the aforementioned topics and see how they match with applied genetic techniques in industry.
Delineating stock management units
While population genetic methods often identify populations and their structure in terms of their evolutionary history alone, the concept of a population is also important for harvested biota as well. Particularly, this applies to the denotation of ‘stocks’ in wild-caught food industries such as fisheries management. Defining stocks helps to limit the amount of harvesting of populations that is sustainable, which is particularly important for species which are distributed across multiple political boundaries (e.g. states or countries).

Monitoring of the health of the various stocks that define a species is important for their long-term sustainability; mismanagement of stocks (either due to a lack of knowledge about stock structure, or directly ignoring catch limits) can dramatically impact the population and broader ecosystem. There are several notable occasions of stock crashes associated with over-depletion of natural populations within fish across the globe, and with the ever-increasing demand for food it will only become more important to analyse and maintain genetic stocks.
Identifying caught species
Higher level taxonomic identification is also important in industry. While this may seem surprising, genetic barcoding methods are important for determining the diversity and numbers of species caught in wider catch methods such as trawling. In many cases, larval or juvenile forms of the caught species are incredibly hard to identify based on morphology, and thus DNA barcoding databases become important for interrogating the impact of these catch methods in the wider ecosystem. Identifying the occurrence and density of different species – both those immediately commercially important and those that are not – is an important evaluator of ecosystem health and sustainability, and might provide early warning signs of ecosystem collapse if important species are missing.

Even at a more direct level, DNA barcoding methods are even used to verify that your own local fishmonger is selling you the correct fish – you’d be surprised how often you might be sold a fish under false pretences. To this end, genetic methods can provide a quality-control method for food being sold to the wider public and reduce the impact of disingenuous marketing practices.
Developing new cultivars
Of course, another aspect of genetic methods in food production that you might be more familiar with is the notion of genetically-modified organisms (GMOs). While GMOs have had considerable debate over the years – some of which academically valid, and others not so much – the ability to improve current food resources is an undoubtable boon for developing nations. Genetic modification of food can take a few different forms based on the available techniques; arguably the most traditional of these is through selective breeding experiments. While perhaps not directly targeted at genes historically, selective breeding of crops for improved yield has occurred for thousands of years. Like with many things, although the ‘adaptive’ genes responsible for these traits are not directly being targeted, the human-induced selective pressure of choosing to reproduce plants (or animals) with desirable traits mimics the process of natural selection well.

Other modification techniques might more directly target the genes of interest, either through more ‘natural’ methods (such as hybridisation of fruit species to provide new seedless varieties) or direct transferal of modified genes through viral or bacterial vectors. These modifications provide new cultivars of food sources to address one issue or another, such as dietary requirements in developing countries or pest resistance to reduce chemical usage, and will seemingly only become more useful with their development.
More to come…
We’ve touched on just a few ways genetic information can be applied to both conservation- and industry-based programs, and demonstrated their shared utility of population genetic theory. Despite their often disparate goals, both industrial and conservation-based groups may apply similar types of information from genetic data to improve their own outcomes. Often, these goals are congruent in driving towards a more sustainable and balanced approach to managing natural resources, an area which is becoming rapidly more relevant as the burden of climate change weighs down upon the Earth.

One thought on “Managing genes in conservation and industry”